Regulation of co-expressed genes
|
 CLARE -- Prediction of regulatory motifs and motif combinations in a set of co-functional enhancers. CLARE -- Prediction of regulatory motifs and motif combinations in a set of co-functional enhancers.
DiRE -- Identification of proximal and Distant Regulatory Elements of co-regulated genes.
SynoR -- Prediction of synonymous regulatory elements in vertebrate genomes.
|
|
Whole genome alignments
|
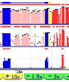 ECR Browser -- Evolutionary conservation of multiple genomes. Identification and sequence analysis of regulatory elements. ECR Browser -- Evolutionary conservation of multiple genomes. Identification and sequence analysis of regulatory elements.
Genome Alignment in ECR Browser -- Align your FASTA nucleotide sequence to a genome of choice.
|
|
Multiple and pairwise sequence alignments
|
 Mulan -- Full multiple sequence alignment. [Interactive conservation profiles, phylogenetic trees, etc.] Mulan -- Full multiple sequence alignment. [Interactive conservation profiles, phylogenetic trees, etc.]
zPicture -- Stacked pairwise and multiple sequence alignment.
|
|
Identification of conserved transcription factor binding sites (cTFBS)
|
 Excluding up to 95% false positive TFBS predictions using sequence conservation as a filter. Excluding up to 95% false positive TFBS predictions using sequence conservation as a filter.
multiTF -- cTFBS in multiple sequence alignments.
rVista 2.0 -- cTFBS in pairwise alignments.
|
|
|



